Biography
Prof Sarah Tabrizi is an award winning scientist who has published over 420 peer-reviewed publications, has been elected a Fellow of the Royal Society, UK Academy of Medical Sciences and US National Academy of Medicine, co-founded the UCL Huntington’s Disease Centre and helped set up the UK All-Party Parliamentary Group for Huntington's disease (HD).
She leads an internationally recognised basic bench science and translational research team focused on understanding mechanism, validating targets and finding disease-modifying therapies for HD. She was PI on the first successful phase 1/2b trial of an antisense oligonucleotide, and currently serves on several SABs advising industry on the development of potential gene targeting and nucleic acid therapies for Huntington’s disease.
Sarah’s research has been recognised by numerous major prizes including the 2019 Yahr Award, 2022 Osler Medal, 2022 HD Society of America Research Award, the 2022 MRC Millennium Medal, the 2023 Arvid Carlsson Award and in 2024 she was elected both to the Fellowship of the Royal Society and the US National Academy of Medicine.
News
Tabrizi Lab
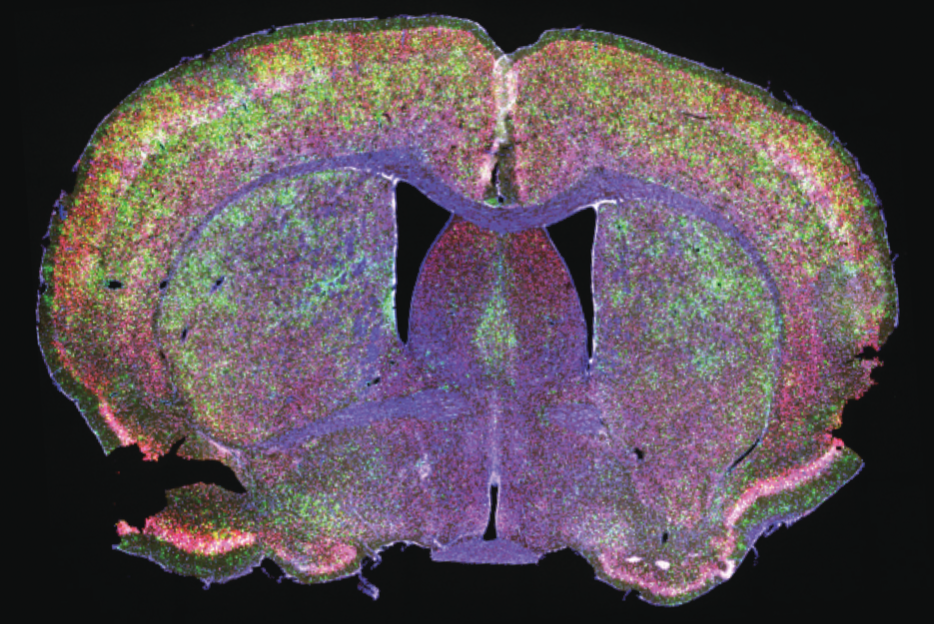
Explore the work of the Tabrizi Lab focused on the modulation of DNA repair and repeat instability in Huntington’s disease and related disorders.