Biography
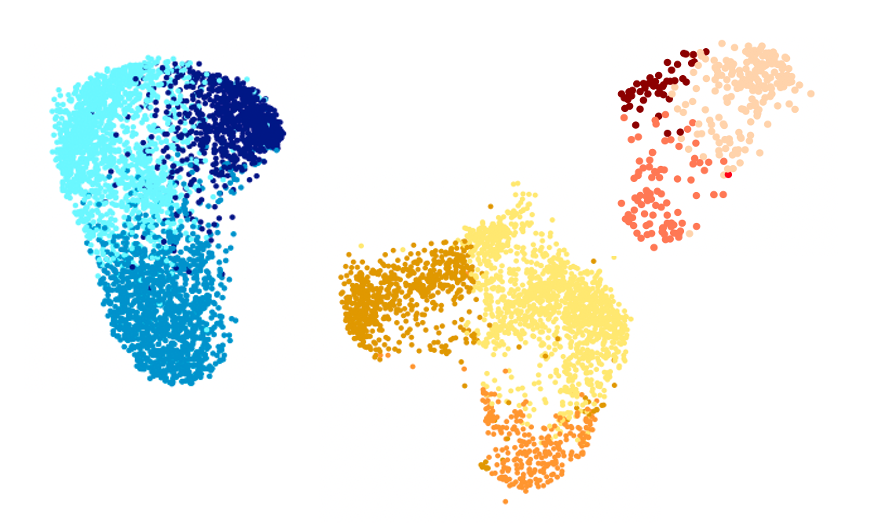
Prof Caleb Webber is combining state-of-the-art stem cell models with bioinformatics techniques to boost our understanding of the biological mechanisms underlying Parkinson’s disease. He is also aiming to identify new risk genes and investigate how these impact on the function of neurons. The team are also using advanced computational methods to investigate how genetic risk factors can influence a varying symptoms presented by patients. They hope to pinpoint key biological pathways that could be targeted with new drugs to prevent or treat Parkinson’s disease– and potentially other diseases with overlapping molecular causes.
In 2022, Prof Webber took on the additional role of UK DRI Director of Data Science. In this role, he is responsible for developing and implementing an Institute-wide strategy to harness the power of data resources and tools in our mission to find new treatments and technologies for dementia.
Webber Lab
Explore the work of the Webber Lab, combining state-of-the-art stem cell models with bioinformatics techniques to boost our understanding of the biological mechanisms underlying Parkinson’s disease