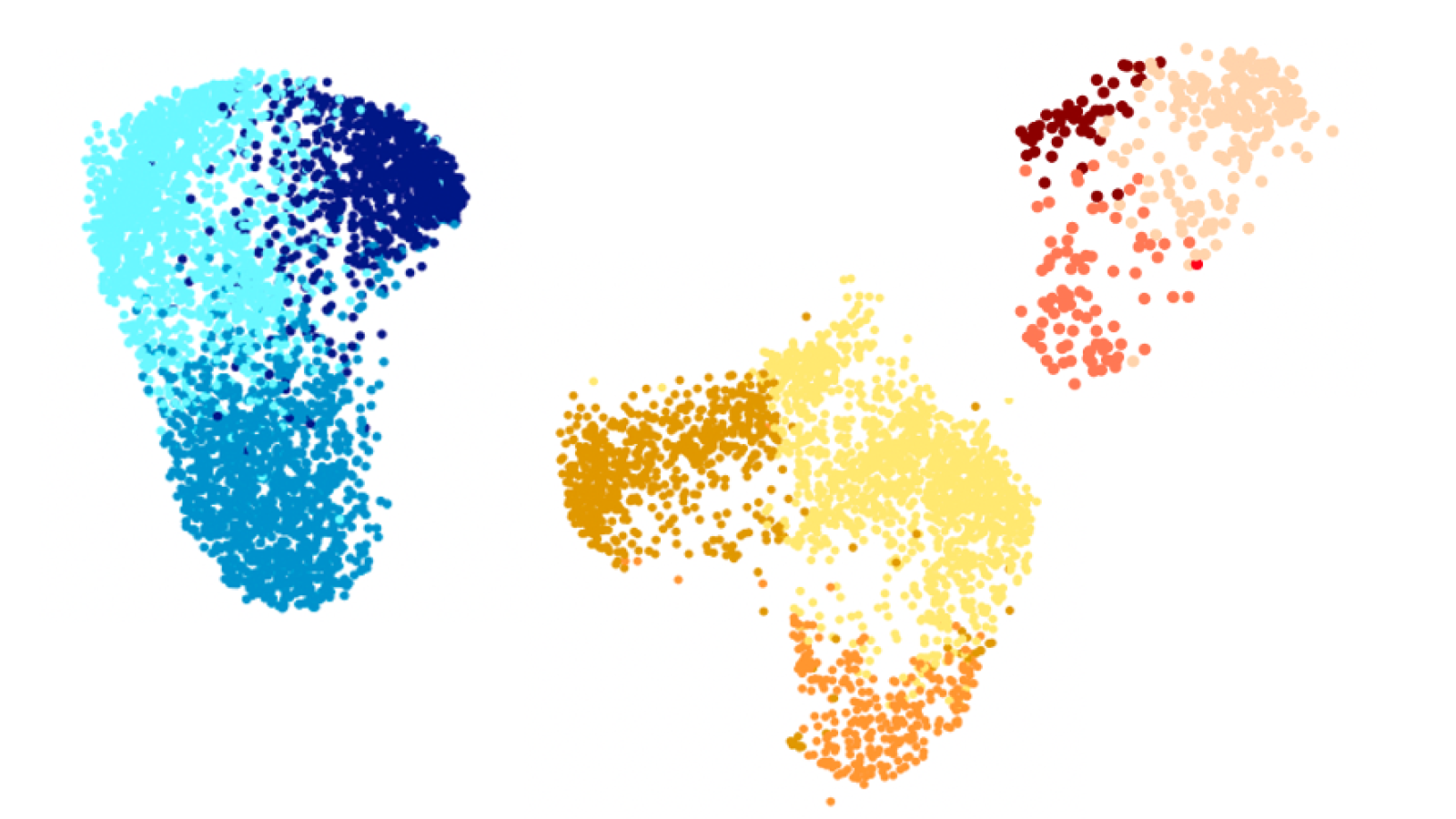
State-of-the-art modelling of Parkinson’s disease
Approximately 145,000 people in the UK are living with Parkinson’s disease. Its symptoms vary widely from person to person, but often include tremors, stiffness and slow movement. These appear when the brain can’t make enough of a chemical called dopamine and gradually get worse over time as more and more dopamine-producing neurons die.
Although the precise causes of Parkinson’s aren’t fully understood, a complex combination of genetic, lifestyle and environmental factors are involved. Using a relatively new approach called genome-wide association studies, scientists have so far identified a number of genetic variations that can influence risk but don’t guarantee that a person will develop the condition.
The Webber Lab is combining state-of-the-art stem cell models with bioinformatics techniques to boost our understanding of the biological mechanisms underlying Parkinson’s disease. He is aiming to identify new risk genes and investigate how these impact on the function of neurons. The team are also using advanced computational methods to investigate how genetic risk factors can influence varying symptoms presented by patients. They hope to pinpoint key biological pathways that could be targeted with new drugs to prevent or treat Parkinson’s disease– and potentially other diseases with overlapping molecular causes.
Latest news

Prof Caleb Webber
Prof Caleb Webber is a Group Leader at the UK DRI at Cardiff, and the UK DRI Director of Data Science. Find out more about his career and expertise on his profile page.

Research summary
Modelling Parkinson’s disease
This UK DRI programme from Prof Caleb Webber is focused on Parkinson’s disease (PD) and split into three key work streams:
(1) Pseudotemporal modelling of disease mechanisms.
Revealing the causal chain of events behind disease-associated variation can yield key insights and offer multiple opportunities to therapeutically intervene. The Webber Lab are applying state-of-the-art single cell pseudotemporal approaches to stem cell models carrying PD-implicated mutations in the genes, SNCA and GBA. The long-term goal is to identify the causal networks and pathways that lead from genetic or environmental insult through to face-valid cellular phenotypes. Their immediate objectives for this stream are to create an iPSC reporter line (LMX1a) that will enable the sorting of neurons without prior fixing, removing heterogeneity from downstream experiments without limiting their study.
(2) Identifying new disease risk genes.
The team is analysing PD genome wide association (GWA) risk loci to identify genes that underlie the risk in these regions. They have made progress in the informatic prioritisation of these genes and will begin rapid modulation of these genes once the LMX1a-reporter line is in place. The long-term objectives of identifying risk mechanisms will involve generating cell-type specific chromatin accessibility maps in iPSC models and nigral brain tissue, and then combining this with fine mapping and CRISPR to identify the risk-modulating genetic variants and their target genes.
(3) Understanding patient-phenotypic heterogeneity.
The team’s new multiple phenotype mixed model approach has identified three axes of variation which have now been re-identified (replicated) in all three deeply-phenotyped PD cohorts in the world. The first two axes alone account for 75% of all clinical observations, and the quantitative traits these axes provide have identified an overlap between the genetic influences of coronary artery disease and Axis 1, and schizophrenia and Axis 2.
Prof Webber’s current research goal is to extend the approach to the longitudinal data, which is already looking very promising. This work is delivering the underlying nature of PD patient phenotypic variation, providing new routes into the influencing biology, and new opportunities to intervene, most rapidly by enabling import of knowledge through shared risk. Furthermore, identifying how PD patients vary is enabling them to devise how deeply-phenotyped cohorts are best phenotyped.
Projects

Key publications
Vacancies
Lab members
Collaborators




Lab funders
Thank you to all those who support the Webber Lab!

