Researchers led by Prof David Klenerman (UK DRI at Cambridge) have designed a new ‘chemical antibody’ that will help them understand the origins of neurological disorders such as Parkinson’s and Alzheimer’s disease.
In many neurodegenerative disorders, proteins become misfolded and form assemblies or ‘aggregates’ that are toxic to brain cells. It is now widely accepted that these larger aggregates are triggered by the accumulation of small, soluble protein bundles called oligomers.
The larger, insoluble aggregates have been intensively studied using conventional methods. However, until now it has not been possible to accurately measure the concentrations of their rare precursor aggregates, or to determine their composition.
In a study published in Nature Chemistry, an international interdisciplinary team of researchers describe how they developed a new molecule that binds to these rare misfolded protein structures, thereby opening the way to a new understanding of how some of the world’s most devastating diseases originate.
As physical chemists we understand how to design, modify and measure the processes of how small molecules interact with biological structures.Prof David KlenermanGroup Leader
A new class of molecule
The story starts with Prof Sonia Gandhi of the Francis Crick Institute and UCL Queen Square Institute of Neurology, who wanted to know how the smaller protein oligomer groups lead to the formation of larger aggregates and eventually to disease. “We have known that these structures are critical to detect,” she said, “But we have never been able to capture them, especially in patients.”
To solve this, Prof Gandhi teamed up with physical and synthetic chemists, including Prof Klenerman and Prof Steven Lee (Professor of Biophysical Chemistry at Cambridge), to design a new class of molecule that would target the oligomers.
“As physical chemists we understand how to design, modify and measure the processes of how small molecules interact with biological structures,” explained Prof Klenerman. He and Prof Lee worked together to design a molecule that would specifically bind to the protein aggregates. “We needed to be able to find a needle in a haystack,” said Prof Lee.
A 'chemical antibody'
The team conceived the idea of a ‘chemical antibody’, which unlike previously designed antibodies could be used to target the structure of protein aggregates rather than their linear sequence. The newly developed Y-shaped small molecule, dubbed “CAP-1”, was designed to target the soluble protein oligomers and bind to them with high affinity, even in complex mixtures such as human cerebrospinal fluid.
Prof Lee explained: “Our new molecule uses the same design principles as found in nature – specifically antibodies – which have an amazing ability to stick to biological structures.”
Next, Prof Tom Snaddon (Department of Chemistry at Indiana University) helped design a way to make the new molecule in the laboratory. Profs Snaddon and Lee already had a long history of working together to create new molecules for tracking and understand neurodegenerative diseases.
“As synthesis scientists, we develop new methods to build molecules,” said Prof Snaddon. “The CAP-1 molecule poses very specific structural challenges, and it took many months of effort before the molecule could be successfully synthesized.”
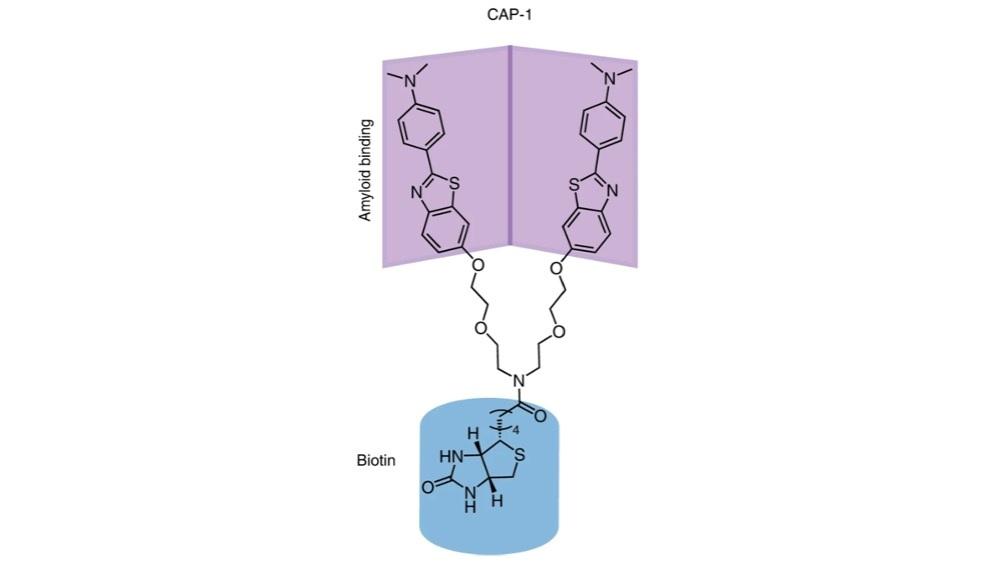
molecule designed to target soluble protein oligomers
Does it work?
Dr Margarida Rodrigues (Faculty of Mathematics at Cambridge), first author of the paper, brought her biochemistry and biophysics background to help determine if the new molecule worked. “Developing a tool capable of isolating all small protein aggregates was exactly the type of exciting and challenging project that would take on,” said Dr Rodrigues. “I was particularly motivated by the ‘out-of-the-box’ design of CAP-1 as compared to the traditional use of antibodies where you can only study one type of protein at a time.”
The researchers spent several years using advanced microscopy methods to investigate how CAP-1 works. They found the new chemical antibody is highly selective to the beta-sheet structure of the protein oligomers, and also has potential advantages in terms of stability and resistance to degradation compared to conventional antibodies. This means that these previously difficult-to-detect protein oligomers can be comprehensively investigated for the first time. "It worked every time! It was so rewarding to ‘see’ CAP-1 capture the aggregates and recover them for further characterisation,” said Dr Rodrigues.
The final step in the process was to find out if the molecule worked in human samples. The group collaborated with Prof Henrik Zetterberg (UK DRI Group Leader at UCL), who has been studying biomarkers for Parkinson’s and Alzheimer’s diseases for over a decade and set up the UK DRI Biomarker Factory which launched earlier this year.
A key experiment was to show that when CAP-1 was used in human cerebrospinal fluid from healthy individuals or patients with Parkinson’s, it could also detect protein aggregates in the fluid, and even enrich them for further studies.
“We anticipate these new tools and methods will be used to detect and characterise the aggregates that initiate and drive the development of neurodegenerative disorders such as Parkinson’s disease,” said Prof Lee.
Read the Alzforum summary of this research.
To find out more about Prof David Klenerman, visit his UK DRI profile. To stay up to date on the latest research news and Institute updates, sign up to receive our monthly newsletter, ‘Inside Eye on UK DRI'.
Source: Yusuf Hamied Department of Chemistry, University of Cambridge
Reference: M. Rodrigues, P. Bhattacharjee, A. Brinkmalm, D.T. Do, C.M. Pearson, S. De, A. Ponjavic, J.A. Varela, K. Kulenkampff, I. Baudrexel, D. Emin, F.S. Ruggeri, J.E. Lee, A.R. Carr, T.P.J. Knowles, H. Zetterberg, T.N. Snaddon, S. Gandhi, S.F. Lee, D. Klenerman, Structure specific amyloid precipitation in biofluids, Nature Chemistry (2022).
Article published: 8 July 2022
Banner image: Shutterstock/ART-ur